Extending the Model in Python
Adding a disease stage
Continuing in ipython/jupyter from the last session, we will now extend the
disease to include an additional, less-infectious, semi-recovering stage,
which will come after I, and be called IR. We do this by inserting a new
stage, named “IR”, at index 2, with beta value 0.2, and progress
value 0.1
>>> lurgy.insert(2, name="IR", beta=0.2, progress=0.1)
>>> print(lurgy)
* Disease: lurgy
* stage: ['E', 'I', 'IR', 'R']
* mapping: ['E', 'I', 'IR', 'R']
* beta: [0, 0.8, 0.2, 0]
* progress: [0.25, 0.25, 0.1, 0]
* too_ill_to_move: [0, 0, 0.0, 0]
* start_symptom: 2
Note
MetaWards is a Python program, so the index is counted from 0. Index 0 is E, index 1 is I and (before this call), index 2 was R. Inserting at index 2 will insert IR between I and R
We can now run the model using metawards.run(). This time we will
set silent to TRUE so that it doesn’t print so much output
to the screen.
>>> results = mw.run(model=home, disease=lurgy,
additional=100, silent=True)
━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━ INFO ━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━
Writing output to directory ./output_s839le7f
━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━
Note
All of the output is written to the (randomly) named output directory
indicated, e.g. for me to output_s839le7f. The full log of the run
is recorded in the file called console.log.bz2 which is in
this directory.
We can now process and plot the results similarly to before, e.g.
>>> df = pd.read_csv(results)
>>> df.plot.line(x="day", y=["S","E","I","IR","R"])
Repeating a run
MetaWards model runs are stochastic, meaning that they use random numbers. While each individual run is reproducible (given the same random number seed and number of processor threads), it is best to run multiple runs so that you can look at averages.
You can perform multiple runs using the repeats argument, e.g.
to perform four runs, you should type;
>>> results = mw.run(model=home, disease=lurgy,
additional=100, silent=True, repeats=4)
If you look at the results, you will that there is a repeat column, which indexes each run with a repeat number, e.g.
>>> df = pd.read_csv(results)
>>> print(df)
fingerprint repeat day date S E I IR R IW
0 REPEAT 1 0 2020-07-22 10000 0 0 0 0 0
1 REPEAT 1 1 2020-07-23 9900 88 12 0 0 1
2 REPEAT 1 2 2020-07-24 9890 74 30 6 0 1
3 REPEAT 1 3 2020-07-25 9864 83 40 13 0 1
4 REPEAT 1 4 2020-07-26 9835 95 48 20 2 1
.. ... ... ... ... ... .. .. .. ... ..
553 REPEAT 4 148 2020-12-17 63 0 0 1 9936 0
554 REPEAT 4 149 2020-12-18 63 0 0 1 9936 0
555 REPEAT 4 150 2020-12-19 63 0 0 1 9936 0
556 REPEAT 4 151 2020-12-20 63 0 0 1 9936 0
557 REPEAT 4 152 2020-12-21 63 0 0 0 9937 0
We can group by repeat and plot using;
>>> import matplotlib.pyplot as plt
>>> df.groupby("repeat").plot.line(
x="day", y=["S","E","I","IR","R"], ax=plt.gca())
You should get a result that looks something like this;
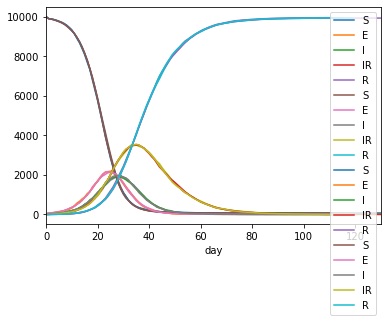
Note
With a bit more pandas you can make this plot a lot prettier ;-)
From this you can see the build-up of individuals in the yellow long recovery (IR) stage.
Adding more wards
Next, we will extend the model by adding more wards. We will model home, work and school, so let’s now add the work and school wards.
>>> work = mw.Ward("work")
>>> school = mw.Ward("school")
We will now add some workers who will make daily, predictable movements from home to work or school.
>>> home.add_workers(7500, destination=work)
>>> home.add_workers(5000, destination=school)
Note
The term worker is very broad in MetaWards. It means any individual that make regular, predictable movements each day. In this case, it refers to workers, teachers and students.
Next we need to combine these individual Ward objects
into a single Wards that represents the entire network.
>>> network = mw.Wards()
>>> network.add(home)
>>> network.add(work)
>>> network.add(school)
Running the model
We can now run the model. In this case, we want to seed the infection in
the home ward, so we need to pass this name into the additional
parameter.
>>> results = mw.run(disease=lurgy, model=network,
additional="1, 100, home")
Note
The format is day number (in this case seed on day 1), then number to seed (seeding 100 infections), then ward name or number (in this case, home)
You will see a lot of output. MetaWards does print a table to confirm the seeding, e.g.
┏━━━━━┳━━━━━━━━━━━━━┳━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━┳━━━━━━━━━━━┓
┃ Day ┃ Demographic ┃ Ward ┃ Number ┃
┃ ┃ ┃ ┃ seeded ┃
┡━━━━━╇━━━━━━━━━━━━━╇━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━╇━━━━━━━━━━━┩
│ 1 │ None │ 1 : WardInfo(name='home', alternate_names=, │ 100 │
│ │ │ code='', alternate_codes=, authority='', │ │
│ │ │ authority_code='', region='', │ │
│ │ │ region_code='') │ │
└─────┴─────────────┴──────────────────────────────────────────────┴───────────┘
The results can be processed and visualised as before, e.g.
>>> df = pd.read_csv(results)
>>> df.plot.line(x="day", y=["S","E","I","IR","R"])
Complete code
The complete Python code for this part of the getting started guide is re-copied below (this continues from the code in the last part);
# extend the disease model
lurgy.insert(2, name="IR", beta=0.2, progress=0.1)
# run the model
results = mw.run(model=home, disease=lurgy,
additional=100, silent=True)
# load and graph the results
df = pd.read_csv(results)
df.plot.line(x="day", y=["S","E","I","IR","R"])
# run multiple repeats
results = mw.run(model=home, disease=lurgy,
additional=100, silent=True, repeats=4)
# load and graph the results
df = pd.read_csv(results)
import matplotlib.pyplot as plt
df.groupby("repeat").plot.line(
x="day", y=["S","E","I","IR","R"], ax=plt.gca())
# add more wards
work = mw.Ward("work")
school = mw.Ward("school")
home.add_workers(7500, destination=work)
home.add_workers(5000, destination=school)
# build the network
network = mw.Wards()
network.add(home)
network.add(work)
network.add(school)
# run the model
results = mw.run(disease=lurgy, model=network,
additional="1, 100, home")
# load the graph the results
df = pd.read_csv(results)
df.plot.line(x="day", y=["S","E","I","IR","R"])